By “group by” we are referring to a process involving one or more of the following steps:
Splitting the data into groups based on some criteria.
Applying a function to each group independently.
Combining the results into a data structure.
Out of these, the split step is the most straightforward. In fact, in many situations we may wish to split the data set into groups and do something with those groups. In the apply step, we might wish to do one of the following:
Aggregation: compute a summary statistic (or statistics) for each group. Some examples:
Transformation: perform some group-specific computations and return a like-indexed object. Some examples:
Filtration: discard some groups, according to a group-wise computation that evaluates True or False. Some examples:
Some combination of the above: GroupBy will examine the results of the apply step and try to return a sensibly combined result if it doesn’t fit into either of the above two categories.
Since the set of object instance methods on pandas data structures are generally rich and expressive, we often simply want to invoke, say, a DataFrame function on each group. The name GroupBy should be quite familiar to those who have used a SQL-based tool (or itertools), in which you can write code like:
SELECT Column1, Column2, mean(Column3), sum(Column4)
FROM SomeTable
GROUP BY Column1, Column2
We aim to make operations like this natural and easy to express using pandas. We’ll address each area of GroupBy functionality then provide some non-trivial examples / use cases.
See the cookbook for some advanced strategies.
Splitting an object into groups
pandas objects can be split on any of their axes. The abstract definition of grouping is to provide a mapping of labels to group names. To create a GroupBy object (more on what the GroupBy object is later), you may do the following:
In [1]: df = pd.DataFrame([('bird', 'Falconiformes', 389.0),
...: ('bird', 'Psittaciformes', 24.0),
...: ('mammal', 'Carnivora', 80.2),
...: ('mammal', 'Primates', np.nan),
...: ('mammal', 'Carnivora', 58)],
...: index=['falcon', 'parrot', 'lion', 'monkey', 'leopard'],
...: columns=('class', 'order', 'max_speed'))
...:
In [2]: df
Out[2]:
class order max_speed
falcon bird Falconiformes 389.0
parrot bird Psittaciformes 24.0
lion mammal Carnivora 80.2
monkey mammal Primates NaN
leopard mammal Carnivora 58.0
# default is axis=0
In [3]: grouped = df.groupby('class')
In [4]: grouped = df.groupby('order', axis='columns')
In [5]: grouped = df.groupby(['class', 'order'])
The mapping can be specified many different ways:
A Python function, to be called on each of the axis labels.
A list or NumPy array of the same length as the selected axis.
A dict or Series, providing a label -> group name mapping.
For DataFrame objects, a string indicating a column to be used to group. Of course df.groupby('A') is just syntactic sugar for df.groupby(df['A']), but it makes life simpler.
For DataFrame objects, a string indicating an index level to be used to group.
A list of any of the above things.
Collectively we refer to the grouping objects as the keys. For example, consider the following DataFrame:
Note
A string passed to groupby may refer to either a column or an index level. If a string matches both a column name and an index level name, a ValueError will be raised.
In [6]: df = pd.DataFrame({'A': ['foo', 'bar', 'foo', 'bar',
...: 'foo', 'bar', 'foo', 'foo'],
...: 'B': ['one', 'one', 'two', 'three',
...: 'two', 'two', 'one', 'three'],
...: 'C': np.random.randn(8),
...: 'D': np.random.randn(8)})
...:
In [7]: df
Out[7]:
A B C D
0 foo one 0.469112 -0.861849
1 bar one -0.282863 -2.104569
2 foo two -1.509059 -0.494929
3 bar three -1.135632 1.071804
4 foo two 1.212112 0.721555
5 bar two -0.173215 -0.706771
6 foo one 0.119209 -1.039575
7 foo three -1.044236 0.271860
On a DataFrame, we obtain a GroupBy object by calling groupby(). We could naturally group by either the A or B columns, or both:
In [8]: grouped = df.groupby('A')
In [9]: grouped = df.groupby(['A', 'B'])
If we also have a MultiIndex on columns A and B, we can group by all but the specified columns
In [10]: df2 = df.set_index(['A', 'B'])
In [11]: grouped = df2.groupby(level=df2.index.names.difference(['B']))
In [12]: grouped.sum()
Out[12]:
C D
A
bar -1.591710 -1.739537
foo -0.752861 -1.402938
These will split the DataFrame on its index (rows). We could also split by the columns:
In [13]: def get_letter_type(letter):
....: if letter.lower() in 'aeiou':
....: return 'vowel'
....: else:
....: return 'consonant'
....:
In [14]: grouped = df.groupby(get_letter_type, axis=1)
pandas Index objects support duplicate values. If a non-unique index is used as the group key in a groupby operation, all values for the same index value will be considered to be in one group and thus the output of aggregation functions will only contain unique index values:
In [15]: lst = [1, 2, 3, 1, 2, 3]
In [16]: s = pd.Series([1, 2, 3, 10, 20, 30], lst)
In [17]: grouped = s.groupby(level=0)
In [18]: grouped.first()
Out[18]:
1 1
2 2
3 3
dtype: int64
In [19]: grouped.last()
Out[19]:
1 10
2 20
3 30
dtype: int64
In [20]: grouped.sum()
Out[20]:
1 11
2 22
3 33
dtype: int64
Note that no splitting occurs until it’s needed. Creating the GroupBy object only verifies that you’ve passed a valid mapping.
Note
Many kinds of complicated data manipulations can be expressed in terms of GroupBy operations (though can’t be guaranteed to be the most efficient). You can get quite creative with the label mapping functions.
GroupBy sorting
By default the group keys are sorted during the groupby operation. You may however pass sort=False for potential speedups:
In [21]: df2 = pd.DataFrame({'X': ['B', 'B', 'A', 'A'], 'Y': [1, 2, 3, 4]})
In [22]: df2.groupby(['X']).sum()
Out[22]:
Y
X
A 7
B 3
In [23]: df2.groupby(['X'], sort=False).sum()
Out[23]:
Y
X
B 3
A 7
Note that groupby will preserve the order in which observations are sorted within each group. For example, the groups created by groupby() below are in the order they appeared in the original DataFrame:
In [24]: df3 = pd.DataFrame({'X': ['A', 'B', 'A', 'B'], 'Y': [1, 4, 3, 2]})
In [25]: df3.groupby(['X']).get_group('A')
Out[25]:
X Y
0 A 1
2 A 3
In [26]: df3.groupby(['X']).get_group('B')
Out[26]:
X Y
1 B 4
3 B 2
GroupBy dropna
By default NA values are excluded from group keys during the groupby operation. However, in case you want to include NA values in group keys, you could pass dropna=False to achieve it.
In [27]: df_list = [[1, 2, 3], [1, None, 4], [2, 1, 3], [1, 2, 2]]
In [28]: df_dropna = pd.DataFrame(df_list, columns=["a", "b", "c"])
In [29]: df_dropna
Out[29]:
a b c
0 1 2.0 3
1 1 NaN 4
2 2 1.0 3
3 1 2.0 2
# Default `dropna` is set to True, which will exclude NaNs in keys
In [30]: df_dropna.groupby(by=["b"], dropna=True).sum()
Out[30]:
a c
b
1.0 2 3
2.0 2 5
# In order to allow NaN in keys, set `dropna` to False
In [31]: df_dropna.groupby(by=["b"], dropna=False).sum()
Out[31]:
a c
b
1.0 2 3
2.0 2 5
NaN 1 4
The default setting of dropna argument is True which means NA are not included in group keys.
GroupBy object attributes
The groups attribute is a dict whose keys are the computed unique groups and corresponding values being the axis labels belonging to each group. In the above example we have:
In [32]: df.groupby('A').groups
Out[32]: {'bar': [1, 3, 5], 'foo': [0, 2, 4, 6, 7]}
In [33]: df.groupby(get_letter_type, axis=1).groups
Out[33]: {'consonant': ['B', 'C', 'D'], 'vowel': ['A']}
Calling the standard Python len function on the GroupBy object just returns the length of the groups dict, so it is largely just a convenience:
In [34]: grouped = df.groupby(['A', 'B'])
In [35]: grouped.groups
Out[35]: {('bar', 'one'): [1], ('bar', 'three'): [3], ('bar', 'two'): [5], ('foo', 'one'): [0, 6], ('foo', 'three'): [7], ('foo', 'two'): [2, 4]}
In [36]: len(grouped)
Out[36]: 6
GroupBy will tab complete column names (and other attributes):
In [37]: df
Out[37]:
height weight gender
2000-01-01 42.849980 157.500553 male
2000-01-02 49.607315 177.340407 male
2000-01-03 56.293531 171.524640 male
2000-01-04 48.421077 144.251986 female
2000-01-05 46.556882 152.526206 male
2000-01-06 68.448851 168.272968 female
2000-01-07 70.757698 136.431469 male
2000-01-08 58.909500 176.499753 female
2000-01-09 76.435631 174.094104 female
2000-01-10 45.306120 177.540920 male
In [38]: gb = df.groupby('gender')
In [39]: gb.<TAB> # noqa: E225, E999
gb.agg gb.boxplot gb.cummin gb.describe gb.filter gb.get_group gb.height gb.last gb.median gb.ngroups gb.plot gb.rank gb.std gb.transform
gb.aggregate gb.count gb.cumprod gb.dtype gb.first gb.groups gb.hist gb.max gb.min gb.nth gb.prod gb.resample gb.sum gb.var
gb.apply gb.cummax gb.cumsum gb.fillna gb.gender gb.head gb.indices gb.mean gb.name gb.ohlc gb.quantile gb.size gb.tail gb.weight
GroupBy with MultiIndex
With hierarchically-indexed data, it’s quite natural to group by one of the levels of the hierarchy.
Let’s create a Series with a two-level MultiIndex.
In [40]: arrays = [['bar', 'bar', 'baz', 'baz', 'foo', 'foo', 'qux', 'qux'],
....: ['one', 'two', 'one', 'two', 'one', 'two', 'one', 'two']]
....:
In [41]: index = pd.MultiIndex.from_arrays(arrays, names=['first', 'second'])
In [42]: s = pd.Series(np.random.randn(8), index=index)
In [43]: s
Out[43]:
first second
bar one -0.919854
two -0.042379
baz one 1.247642
two -0.009920
foo one 0.290213
two 0.495767
qux one 0.362949
two 1.548106
dtype: float64
We can then group by one of the levels in s.
In [44]: grouped = s.groupby(level=0)
In [45]: grouped.sum()
Out[45]:
first
bar -0.962232
baz 1.237723
foo 0.785980
qux 1.911055
dtype: float64
If the MultiIndex has names specified, these can be passed instead of the level number:
In [46]: s.groupby(level='second').sum()
Out[46]:
second
one 0.980950
two 1.991575
dtype: float64
The aggregation functions such as sum will take the level parameter directly. Additionally, the resulting index will be named according to the chosen level:
In [47]: s.sum(level='second')
Out[47]:
second
one 0.980950
two 1.991575
dtype: float64
Grouping with multiple levels is supported.
In [48]: s
Out[48]:
first second third
bar doo one -1.131345
two -0.089329
baz bee one 0.337863
two -0.945867
foo bop one -0.932132
two 1.956030
qux bop one 0.017587
two -0.016692
dtype: float64
In [49]: s.groupby(level=['first', 'second']).sum()
Out[49]:
first second
bar doo -1.220674
baz bee -0.608004
foo bop 1.023898
qux bop 0.000895
dtype: float64
Index level names may be supplied as keys.
In [50]: s.groupby(['first', 'second']).sum()
Out[50]:
first second
bar doo -1.220674
baz bee -0.608004
foo bop 1.023898
qux bop 0.000895
dtype: float64
More on the sum function and aggregation later.
Grouping DataFrame with Index levels and columns
A DataFrame may be grouped by a combination of columns and index levels by specifying the column names as strings and the index levels as pd.Grouper objects.
In [51]: arrays = [['bar', 'bar', 'baz', 'baz', 'foo', 'foo', 'qux', 'qux'],
....: ['one', 'two', 'one', 'two', 'one', 'two', 'one', 'two']]
....:
In [52]: index = pd.MultiIndex.from_arrays(arrays, names=['first', 'second'])
In [53]: df = pd.DataFrame({'A': [1, 1, 1, 1, 2, 2, 3, 3],
....: 'B': np.arange(8)},
....: index=index)
....:
In [54]: df
Out[54]:
A B
first second
bar one 1 0
two 1 1
baz one 1 2
two 1 3
foo one 2 4
two 2 5
qux one 3 6
two 3 7
The following example groups df by the second index level and the A column.
In [55]: df.groupby([pd.Grouper(level=1), 'A']).sum()
Out[55]:
B
second A
one 1 2
2 4
3 6
two 1 4
2 5
3 7
Index levels may also be specified by name.
In [56]: df.groupby([pd.Grouper(level='second'), 'A']).sum()
Out[56]:
B
second A
one 1 2
2 4
3 6
two 1 4
2 5
3 7
Index level names may be specified as keys directly to groupby.
In [57]: df.groupby(['second', 'A']).sum()
Out[57]:
B
second A
one 1 2
2 4
3 6
two 1 4
2 5
3 7
DataFrame column selection in GroupBy
Once you have created the GroupBy object from a DataFrame, you might want to do something different for each of the columns. Thus, using [] similar to getting a column from a DataFrame, you can do:
In [58]: grouped = df.groupby(['A'])
In [59]: grouped_C = grouped['C']
In [60]: grouped_D = grouped['D']
This is mainly syntactic sugar for the alternative and much more verbose:
In [61]: df['C'].groupby(df['A'])
Out[61]: <pandas.core.groupby.generic.SeriesGroupBy object at 0x7f9da1c54d00>
Additionally this method avoids recomputing the internal grouping information derived from the passed key.
Iterating through groups
With the GroupBy object in hand, iterating through the grouped data is very natural and functions similarly to itertools.groupby():
In [62]: grouped = df.groupby('A')
In [63]: for name, group in grouped:
....: print(name)
....: print(group)
....:
bar
A B C D
1 bar one 0.254161 1.511763
3 bar three 0.215897 -0.990582
5 bar two -0.077118 1.211526
foo
A B C D
0 foo one -0.575247 1.346061
2 foo two -1.143704 1.627081
4 foo two 1.193555 -0.441652
6 foo one -0.408530 0.268520
7 foo three -0.862495 0.024580
In the case of grouping by multiple keys, the group name will be a tuple:
In [64]: for name, group in df.groupby(['A', 'B']):
....: print(name)
....: print(group)
....:
('bar', 'one')
A B C D
1 bar one 0.254161 1.511763
('bar', 'three')
A B C D
3 bar three 0.215897 -0.990582
('bar', 'two')
A B C D
5 bar two -0.077118 1.211526
('foo', 'one')
A B C D
0 foo one -0.575247 1.346061
6 foo one -0.408530 0.268520
('foo', 'three')
A B C D
7 foo three -0.862495 0.02458
('foo', 'two')
A B C D
2 foo two -1.143704 1.627081
4 foo two 1.193555 -0.441652
See Iterating through groups.
Selecting a group
A single group can be selected using get_group():
In [65]: grouped.get_group('bar')
Out[65]:
A B C D
1 bar one 0.254161 1.511763
3 bar three 0.215897 -0.990582
5 bar two -0.077118 1.211526
Or for an object grouped on multiple columns:
In [66]: df.groupby(['A', 'B']).get_group(('bar', 'one'))
Out[66]:
A B C D
1 bar one 0.254161 1.511763
Aggregation
Once the GroupBy object has been created, several methods are available to perform a computation on the grouped data. These operations are similar to the aggregating API, window functions API, and resample API.
An obvious one is aggregation via the aggregate() or equivalently agg() method:
In [67]: grouped = df.groupby('A')
In [68]: grouped.aggregate(np.sum)
Out[68]:
C D
A
bar 0.392940 1.732707
foo -1.796421 2.824590
In [69]: grouped = df.groupby(['A', 'B'])
In [70]: grouped.aggregate(np.sum)
Out[70]:
C D
A B
bar one 0.254161 1.511763
three 0.215897 -0.990582
two -0.077118 1.211526
foo one -0.983776 1.614581
three -0.862495 0.024580
two 0.049851 1.185429
As you can see, the result of the aggregation will have the group names as the new index along the grouped axis. In the case of multiple keys, the result is a MultiIndex by default, though this can be changed by using the as_index option:
In [71]: grouped = df.groupby(['A', 'B'], as_index=False)
In [72]: grouped.aggregate(np.sum)
Out[72]:
A B C D
0 bar one 0.254161 1.511763
1 bar three 0.215897 -0.990582
2 bar two -0.077118 1.211526
3 foo one -0.983776 1.614581
4 foo three -0.862495 0.024580
5 foo two 0.049851 1.185429
In [73]: df.groupby('A', as_index=False).sum()
Out[73]:
A C D
0 bar 0.392940 1.732707
1 foo -1.796421 2.824590
Note that you could use the reset_index DataFrame function to achieve the same result as the column names are stored in the resulting MultiIndex:
In [74]: df.groupby(['A', 'B']).sum().reset_index()
Out[74]:
A B C D
0 bar one 0.254161 1.511763
1 bar three 0.215897 -0.990582
2 bar two -0.077118 1.211526
3 foo one -0.983776 1.614581
4 foo three -0.862495 0.024580
5 foo two 0.049851 1.185429
Another simple aggregation example is to compute the size of each group. This is included in GroupBy as the size method. It returns a Series whose index are the group names and whose values are the sizes of each group.
In [75]: grouped.size()
Out[75]:
A B size
0 bar one 1
1 bar three 1
2 bar two 1
3 foo one 2
4 foo three 1
5 foo two 2
In [76]: grouped.describe()
Out[76]:
C ... D
count mean std min 25% 50% 75% ... mean std min 25% 50% 75% max
0 1.0 0.254161 NaN 0.254161 0.254161 0.254161 0.254161 ... 1.511763 NaN 1.511763 1.511763 1.511763 1.511763 1.511763
1 1.0 0.215897 NaN 0.215897 0.215897 0.215897 0.215897 ... -0.990582 NaN -0.990582 -0.990582 -0.990582 -0.990582 -0.990582
2 1.0 -0.077118 NaN -0.077118 -0.077118 -0.077118 -0.077118 ... 1.211526 NaN 1.211526 1.211526 1.211526 1.211526 1.211526
3 2.0 -0.491888 0.117887 -0.575247 -0.533567 -0.491888 -0.450209 ... 0.807291 0.761937 0.268520 0.537905 0.807291 1.076676 1.346061
4 1.0 -0.862495 NaN -0.862495 -0.862495 -0.862495 -0.862495 ... 0.024580 NaN 0.024580 0.024580 0.024580 0.024580 0.024580
5 2.0 0.024925 1.652692 -1.143704 -0.559389 0.024925 0.609240 ... 0.592714 1.462816 -0.441652 0.075531 0.592714 1.109898 1.627081
[6 rows x 16 columns]
Note
Aggregation functions will not return the groups that you are aggregating over if they are named columns, when as_index=True, the default. The grouped columns will be the indices of the returned object.
Passing as_index=False will return the groups that you are aggregating over, if they are named columns.
Aggregating functions are the ones that reduce the dimension of the returned objects. Some common aggregating functions are tabulated below:
Function | Description |
|---|
mean()
| Compute mean of groups |
sum()
| Compute sum of group values |
size()
| Compute group sizes |
count()
| Compute count of group |
std()
| Standard deviation of groups |
var()
| Compute variance of groups |
sem()
| Standard error of the mean of groups |
describe()
| Generates descriptive statistics |
first()
| Compute first of group values |
last()
| Compute last of group values |
nth()
| Take nth value, or a subset if n is a list |
min()
| Compute min of group values |
max()
| Compute max of group values |
The aggregating functions above will exclude NA values. Any function which reduces a Series to a scalar value is an aggregation function and will work, a trivial example is df.groupby('A').agg(lambda ser: 1). Note that nth() can act as a reducer or a filter, see here.
Applying multiple functions at once
With grouped Series you can also pass a list or dict of functions to do aggregation with, outputting a DataFrame:
In [77]: grouped = df.groupby('A')
In [78]: grouped['C'].agg([np.sum, np.mean, np.std])
Out[78]:
sum mean std
A
bar 0.392940 0.130980 0.181231
foo -1.796421 -0.359284 0.912265
On a grouped DataFrame, you can pass a list of functions to apply to each column, which produces an aggregated result with a hierarchical index:
In [79]: grouped.agg([np.sum, np.mean, np.std])
Out[79]:
C D
sum mean std sum mean std
A
bar 0.392940 0.130980 0.181231 1.732707 0.577569 1.366330
foo -1.796421 -0.359284 0.912265 2.824590 0.564918 0.884785
The resulting aggregations are named for the functions themselves. If you need to rename, then you can add in a chained operation for a Series like this:
In [80]: (grouped['C'].agg([np.sum, np.mean, np.std])
....: .rename(columns={'sum': 'foo',
....: 'mean': 'bar',
....: 'std': 'baz'}))
....:
Out[80]:
foo bar baz
A
bar 0.392940 0.130980 0.181231
foo -1.796421 -0.359284 0.912265
For a grouped DataFrame, you can rename in a similar manner:
In [81]: (grouped.agg([np.sum, np.mean, np.std])
....: .rename(columns={'sum': 'foo',
....: 'mean': 'bar',
....: 'std': 'baz'}))
....:
Out[81]:
C D
foo bar baz foo bar baz
A
bar 0.392940 0.130980 0.181231 1.732707 0.577569 1.366330
foo -1.796421 -0.359284 0.912265 2.824590 0.564918 0.884785
Note
In general, the output column names should be unique. You can’t apply the same function (or two functions with the same name) to the same column.
In [82]: grouped['C'].agg(['sum', 'sum'])
Out[82]:
sum sum
A
bar 0.392940 0.392940
foo -1.796421 -1.796421
Pandas does allow you to provide multiple lambdas. In this case, pandas will mangle the name of the (nameless) lambda functions, appending _<i> to each subsequent lambda.
In [83]: grouped['C'].agg([lambda x: x.max() - x.min(),
....: lambda x: x.median() - x.mean()])
....:
Out[83]:
<lambda_0> <lambda_1>
A
bar 0.331279 0.084917
foo 2.337259 -0.215962
Named aggregation
To support column-specific aggregation with control over the output column names, pandas accepts the special syntax in GroupBy.agg(), known as “named aggregation”, where
The keywords are the output column names
The values are tuples whose first element is the column to select and the second element is the aggregation to apply to that column. Pandas provides the pandas.NamedAgg namedtuple with the fields ['column', 'aggfunc'] to make it clearer what the arguments are. As usual, the aggregation can be a callable or a string alias.
In [84]: animals = pd.DataFrame({'kind': ['cat', 'dog', 'cat', 'dog'],
....: 'height': [9.1, 6.0, 9.5, 34.0],
....: 'weight': [7.9, 7.5, 9.9, 198.0]})
....:
In [85]: animals
Out[85]:
kind height weight
0 cat 9.1 7.9
1 dog 6.0 7.5
2 cat 9.5 9.9
3 dog 34.0 198.0
In [86]: animals.groupby("kind").agg(
....: min_height=pd.NamedAgg(column='height', aggfunc='min'),
....: max_height=pd.NamedAgg(column='height', aggfunc='max'),
....: average_weight=pd.NamedAgg(column='weight', aggfunc=np.mean),
....: )
....:
Out[86]:
min_height max_height average_weight
kind
cat 9.1 9.5 8.90
dog 6.0 34.0 102.75
pandas.NamedAgg is just a namedtuple. Plain tuples are allowed as well.
In [87]: animals.groupby("kind").agg(
....: min_height=('height', 'min'),
....: max_height=('height', 'max'),
....: average_weight=('weight', np.mean),
....: )
....:
Out[87]:
min_height max_height average_weight
kind
cat 9.1 9.5 8.90
dog 6.0 34.0 102.75
If your desired output column names are not valid python keywords, construct a dictionary and unpack the keyword arguments
In [88]: animals.groupby("kind").agg(**{
....: 'total weight': pd.NamedAgg(column='weight', aggfunc=sum),
....: })
....:
Out[88]:
total weight
kind
cat 17.8
dog 205.5
Additional keyword arguments are not passed through to the aggregation functions. Only pairs of (column, aggfunc) should be passed as **kwargs. If your aggregation functions requires additional arguments, partially apply them with functools.partial().
Note
For Python 3.5 and earlier, the order of **kwargs in a functions was not preserved. This means that the output column ordering would not be consistent. To ensure consistent ordering, the keys (and so output columns) will always be sorted for Python 3.5.
Named aggregation is also valid for Series groupby aggregations. In this case there’s no column selection, so the values are just the functions.
In [89]: animals.groupby("kind").height.agg(
....: min_height='min',
....: max_height='max',
....: )
....:
Out[89]:
min_height max_height
kind
cat 9.1 9.5
dog 6.0 34.0
Applying different functions to DataFrame columns
By passing a dict to aggregate you can apply a different aggregation to the columns of a DataFrame:
In [90]: grouped.agg({'C': np.sum,
....: 'D': lambda x: np.std(x, ddof=1)})
....:
Out[90]:
C D
A
bar 0.392940 1.366330
foo -1.796421 0.884785
The function names can also be strings. In order for a string to be valid it must be either implemented on GroupBy or available via dispatching:
In [91]: grouped.agg({'C': 'sum', 'D': 'std'})
Out[91]:
C D
A
bar 0.392940 1.366330
foo -1.796421 0.884785
Cython-optimized aggregation functions
Some common aggregations, currently only sum, mean, std, and sem, have optimized Cython implementations:
In [92]: df.groupby('A').sum()
Out[92]:
C D
A
bar 0.392940 1.732707
foo -1.796421 2.824590
In [93]: df.groupby(['A', 'B']).mean()
Out[93]:
C D
A B
bar one 0.254161 1.511763
three 0.215897 -0.990582
two -0.077118 1.211526
foo one -0.491888 0.807291
three -0.862495 0.024580
two 0.024925 0.592714
Of course sum and mean are implemented on pandas objects, so the above code would work even without the special versions via dispatching (see below).
Filtration
The filter method returns a subset of the original object. Suppose we want to take only elements that belong to groups with a group sum greater than 2.
In [132]: sf = pd.Series([1, 1, 2, 3, 3, 3])
In [133]: sf.groupby(sf).filter(lambda x: x.sum() > 2)
Out[133]:
3 3
4 3
5 3
dtype: int64
The argument of filter must be a function that, applied to the group as a whole, returns True or False.
Another useful operation is filtering out elements that belong to groups with only a couple members.
In [134]: dff = pd.DataFrame({'A': np.arange(8), 'B': list('aabbbbcc')})
In [135]: dff.groupby('B').filter(lambda x: len(x) > 2)
Out[135]:
A B
2 2 b
3 3 b
4 4 b
5 5 b
Alternatively, instead of dropping the offending groups, we can return a like-indexed objects where the groups that do not pass the filter are filled with NaNs.
In [136]: dff.groupby('B').filter(lambda x: len(x) > 2, dropna=False)
Out[136]:
A B
0 NaN NaN
1 NaN NaN
2 2.0 b
3 3.0 b
4 4.0 b
5 5.0 b
6 NaN NaN
7 NaN NaN
For DataFrames with multiple columns, filters should explicitly specify a column as the filter criterion.
In [137]: dff['C'] = np.arange(8)
In [138]: dff.groupby('B').filter(lambda x: len(x['C']) > 2)
Out[138]:
A B C
2 2 b 2
3 3 b 3
4 4 b 4
5 5 b 5
Note
Some functions when applied to a groupby object will act as a filter on the input, returning a reduced shape of the original (and potentially eliminating groups), but with the index unchanged. Passing as_index=False will not affect these transformation methods.
For example: head, tail.
In [139]: dff.groupby('B').head(2)
Out[139]:
A B C
0 0 a 0
1 1 a 1
2 2 b 2
3 3 b 3
6 6 c 6
7 7 c 7
Dispatching to instance methods
When doing an aggregation or transformation, you might just want to call an instance method on each data group. This is pretty easy to do by passing lambda functions:
In [140]: grouped = df.groupby('A')
In [141]: grouped.agg(lambda x: x.std())
Out[141]:
C D
A
bar 0.181231 1.366330
foo 0.912265 0.884785
But, it’s rather verbose and can be untidy if you need to pass additional arguments. Using a bit of metaprogramming cleverness, GroupBy now has the ability to “dispatch” method calls to the groups:
In [142]: grouped.std()
Out[142]:
C D
A
bar 0.181231 1.366330
foo 0.912265 0.884785
What is actually happening here is that a function wrapper is being generated. When invoked, it takes any passed arguments and invokes the function with any arguments on each group (in the above example, the std function). The results are then combined together much in the style of agg and transform (it actually uses apply to infer the gluing, documented next). This enables some operations to be carried out rather succinctly:
In [143]: tsdf = pd.DataFrame(np.random.randn(1000, 3),
.....: index=pd.date_range('1/1/2000', periods=1000),
.....: columns=['A', 'B', 'C'])
.....:
In [144]: tsdf.iloc[::2] = np.nan
In [145]: grouped = tsdf.groupby(lambda x: x.year)
In [146]: grouped.fillna(method='pad')
Out[146]:
A B C
2000-01-01 NaN NaN NaN
2000-01-02 -0.353501 -0.080957 -0.876864
2000-01-03 -0.353501 -0.080957 -0.876864
2000-01-04 0.050976 0.044273 -0.559849
2000-01-05 0.050976 0.044273 -0.559849
... ... ... ...
2002-09-22 0.005011 0.053897 -1.026922
2002-09-23 0.005011 0.053897 -1.026922
2002-09-24 -0.456542 -1.849051 1.559856
2002-09-25 -0.456542 -1.849051 1.559856
2002-09-26 1.123162 0.354660 1.128135
[1000 rows x 3 columns]
In this example, we chopped the collection of time series into yearly chunks then independently called fillna on the groups.
The nlargest and nsmallest methods work on Series style groupbys:
In [147]: s = pd.Series([9, 8, 7, 5, 19, 1, 4.2, 3.3])
In [148]: g = pd.Series(list('abababab'))
In [149]: gb = s.groupby(g)
In [150]: gb.nlargest(3)
Out[150]:
a 4 19.0
0 9.0
2 7.0
b 1 8.0
3 5.0
7 3.3
dtype: float64
In [151]: gb.nsmallest(3)
Out[151]:
a 6 4.2
2 7.0
0 9.0
b 5 1.0
7 3.3
3 5.0
dtype: float64
Flexible apply
Some operations on the grouped data might not fit into either the aggregate or transform categories. Or, you may simply want GroupBy to infer how to combine the results. For these, use the apply function, which can be substituted for both aggregate and transform in many standard use cases. However, apply can handle some exceptional use cases, for example:
In [152]: df
Out[152]:
A B C D
0 foo one -0.575247 1.346061
1 bar one 0.254161 1.511763
2 foo two -1.143704 1.627081
3 bar three 0.215897 -0.990582
4 foo two 1.193555 -0.441652
5 bar two -0.077118 1.211526
6 foo one -0.408530 0.268520
7 foo three -0.862495 0.024580
In [153]: grouped = df.groupby('A')
# could also just call .describe()
In [154]: grouped['C'].apply(lambda x: x.describe())
Out[154]:
A
bar count 3.000000
mean 0.130980
std 0.181231
min -0.077118
25% 0.069390
...
foo min -1.143704
25% -0.862495
50% -0.575247
75% -0.408530
max 1.193555
Name: C, Length: 16, dtype: float64
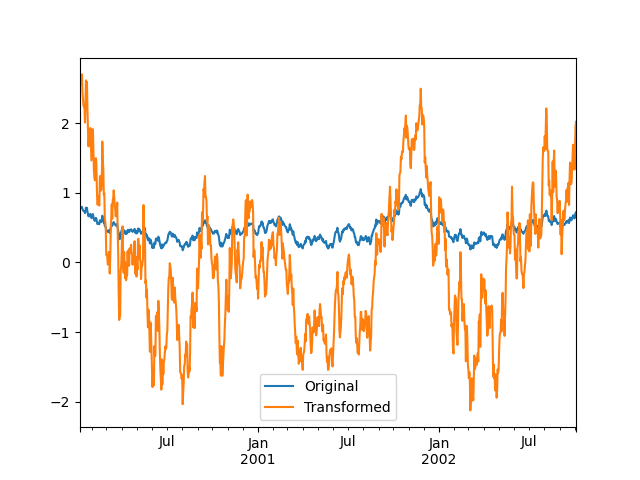
The dimension of the returned result can also change:
In [155]: grouped = df.groupby('A')['C']
In [156]: def f(group):
.....: return pd.DataFrame({'original': group,
.....: 'demeaned': group - group.mean()})
.....:
In [157]: grouped.apply(f)
Out[157]:
original demeaned
0 -0.575247 -0.215962
1 0.254161 0.123181
2 -1.143704 -0.784420
3 0.215897 0.084917
4 1.193555 1.552839
5 -0.077118 -0.208098
6 -0.408530 -0.049245
7 -0.862495 -0.503211
apply on a Series can operate on a returned value from the applied function, that is itself a series, and possibly upcast the result to a DataFrame:
In [158]: def f(x):
.....: return pd.Series([x, x ** 2], index=['x', 'x^2'])
.....:
In [159]: s = pd.Series(np.random.rand(5))
In [160]: s
Out[160]:
0 0.321438
1 0.493496
2 0.139505
3 0.910103
4 0.194158
dtype: float64
In [161]: s.apply(f)
Out[161]:
x x^2
0 0.321438 0.103323
1 0.493496 0.243538
2 0.139505 0.019462
3 0.910103 0.828287
4 0.194158 0.037697
Note
apply can act as a reducer, transformer, or filter function, depending on exactly what is passed to it. So depending on the path taken, and exactly what you are grouping. Thus the grouped columns(s) may be included in the output as well as set the indices.
Numba Accelerated Routines
If Numba is installed as an optional dependency, the transform and aggregate methods support engine='numba' and engine_kwargs arguments. The engine_kwargs argument is a dictionary of keyword arguments that will be passed into the numba.jit decorator. These keyword arguments will be applied to the passed function. Currently only nogil, nopython, and parallel are supported, and their default values are set to False, True and False respectively.
The function signature must start with values, index exactly as the data belonging to each group will be passed into values, and the group index will be passed into index.
Warning
When using engine='numba', there will be no “fall back” behavior internally. The group data and group index will be passed as numpy arrays to the JITed user defined function, and no alternative execution attempts will be tried.
Note
In terms of performance, the first time a function is run using the Numba engine will be slow as Numba will have some function compilation overhead. However, the compiled functions are cached, and subsequent calls will be fast. In general, the Numba engine is performant with a larger amount of data points (e.g. 1+ million).
In [1]: N = 10 ** 3
In [2]: data = {0: [str(i) for i in range(100)] * N, 1: list(range(100)) * N}
In [3]: df = pd.DataFrame(data, columns=[0, 1])
In [4]: def f_numba(values, index):
...: total = 0
...: for i, value in enumerate(values):
...: if i % 2:
...: total += value + 5
...: else:
...: total += value * 2
...: return total
...:
In [5]: def f_cython(values):
...: total = 0
...: for i, value in enumerate(values):
...: if i % 2:
...: total += value + 5
...: else:
...: total += value * 2
...: return total
...:
In [6]: groupby = df.groupby(0)
# Run the first time, compilation time will affect performance
In [7]: %timeit -r 1 -n 1 groupby.aggregate(f_numba, engine='numba') # noqa: E225
2.14 s ± 0 ns per loop (mean ± std. dev. of 1 run, 1 loop each)
# Function is cached and performance will improve
In [8]: %timeit groupby.aggregate(f_numba, engine='numba')
4.93 ms ± 32.3 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
In [9]: %timeit groupby.aggregate(f_cython, engine='cython')
18.6 ms ± 84.8 µs per loop (mean ± std. dev. of 7 runs, 100 loops each)
Other useful features
Automatic exclusion of “nuisance” columns
Again consider the example DataFrame we’ve been looking at:
In [162]: df
Out[162]:
A B C D
0 foo one -0.575247 1.346061
1 bar one 0.254161 1.511763
2 foo two -1.143704 1.627081
3 bar three 0.215897 -0.990582
4 foo two 1.193555 -0.441652
5 bar two -0.077118 1.211526
6 foo one -0.408530 0.268520
7 foo three -0.862495 0.024580
Suppose we wish to compute the standard deviation grouped by the A column. There is a slight problem, namely that we don’t care about the data in column B. We refer to this as a “nuisance” column. If the passed aggregation function can’t be applied to some columns, the troublesome columns will be (silently) dropped. Thus, this does not pose any problems:
In [163]: df.groupby('A').std()
Out[163]:
C D
A
bar 0.181231 1.366330
foo 0.912265 0.884785
Note that df.groupby('A').colname.std(). is more efficient than df.groupby('A').std().colname, so if the result of an aggregation function is only interesting over one column (here colname), it may be filtered before applying the aggregation function.
Note
Any object column, also if it contains numerical values such as Decimal objects, is considered as a “nuisance” columns. They are excluded from aggregate functions automatically in groupby.
If you do wish to include decimal or object columns in an aggregation with other non-nuisance data types, you must do so explicitly.
In [164]: from decimal import Decimal
In [165]: df_dec = pd.DataFrame(
.....: {'id': [1, 2, 1, 2],
.....: 'int_column': [1, 2, 3, 4],
.....: 'dec_column': [Decimal('0.50'), Decimal('0.15'),
.....: Decimal('0.25'), Decimal('0.40')]
.....: }
.....: )
.....:
# Decimal columns can be sum'd explicitly by themselves...
In [166]: df_dec.groupby(['id'])[['dec_column']].sum()
Out[166]:
dec_column
id
1 0.75
2 0.55
# ...but cannot be combined with standard data types or they will be excluded
In [167]: df_dec.groupby(['id'])[['int_column', 'dec_column']].sum()
Out[167]:
int_column
id
1 4
2 6
# Use .agg function to aggregate over standard and "nuisance" data types
# at the same time
In [168]: df_dec.groupby(['id']).agg({'int_column': 'sum', 'dec_column': 'sum'})
Out[168]:
int_column dec_column
id
1 4 0.75
2 6 0.55
Handling of (un)observed Categorical values
When using a Categorical grouper (as a single grouper, or as part of multiple groupers), the observed keyword controls whether to return a cartesian product of all possible groupers values (observed=False) or only those that are observed groupers (observed=True).
Show all values:
In [169]: pd.Series([1, 1, 1]).groupby(pd.Categorical(['a', 'a', 'a'],
.....: categories=['a', 'b']),
.....: observed=False).count()
.....:
Out[169]:
a 3
b 0
dtype: int64
Show only the observed values:
In [170]: pd.Series([1, 1, 1]).groupby(pd.Categorical(['a', 'a', 'a'],
.....: categories=['a', 'b']),
.....: observed=True).count()
.....:
Out[170]:
a 3
dtype: int64
The returned dtype of the grouped will always include all of the categories that were grouped.
In [171]: s = pd.Series([1, 1, 1]).groupby(pd.Categorical(['a', 'a', 'a'],
.....: categories=['a', 'b']),
.....: observed=False).count()
.....:
In [172]: s.index.dtype
Out[172]: CategoricalDtype(categories=['a', 'b'], ordered=False)
NA and NaT group handling
If there are any NaN or NaT values in the grouping key, these will be automatically excluded. In other words, there will never be an “NA group” or “NaT group”. This was not the case in older versions of pandas, but users were generally discarding the NA group anyway (and supporting it was an implementation headache).
Grouping with ordered factors
Categorical variables represented as instance of pandas’s Categorical class can be used as group keys. If so, the order of the levels will be preserved:
In [173]: data = pd.Series(np.random.randn(100))
In [174]: factor = pd.qcut(data, [0, .25, .5, .75, 1.])
In [175]: data.groupby(factor).mean()
Out[175]:
(-2.645, -0.523] -1.362896
(-0.523, 0.0296] -0.260266
(0.0296, 0.654] 0.361802
(0.654, 2.21] 1.073801
dtype: float64
Grouping with a grouper specification
You may need to specify a bit more data to properly group. You can use the pd.Grouper to provide this local control.
In [176]: import datetime
In [177]: df = pd.DataFrame({'Branch': 'A A A A A A A B'.split(),
.....: 'Buyer': 'Carl Mark Carl Carl Joe Joe Joe Carl'.split(),
.....: 'Quantity': [1, 3, 5, 1, 8, 1, 9, 3],
.....: 'Date': [
.....: datetime.datetime(2013, 1, 1, 13, 0),
.....: datetime.datetime(2013, 1, 1, 13, 5),
.....: datetime.datetime(2013, 10, 1, 20, 0),
.....: datetime.datetime(2013, 10, 2, 10, 0),
.....: datetime.datetime(2013, 10, 1, 20, 0),
.....: datetime.datetime(2013, 10, 2, 10, 0),
.....: datetime.datetime(2013, 12, 2, 12, 0),
.....: datetime.datetime(2013, 12, 2, 14, 0)]
.....: })
.....:
In [178]: df
Out[178]:
Branch Buyer Quantity Date
0 A Carl 1 2013-01-01 13:00:00
1 A Mark 3 2013-01-01 13:05:00
2 A Carl 5 2013-10-01 20:00:00
3 A Carl 1 2013-10-02 10:00:00
4 A Joe 8 2013-10-01 20:00:00
5 A Joe 1 2013-10-02 10:00:00
6 A Joe 9 2013-12-02 12:00:00
7 B Carl 3 2013-12-02 14:00:00
Groupby a specific column with the desired frequency. This is like resampling.
In [179]: df.groupby([pd.Grouper(freq='1M', key='Date'), 'Buyer']).sum()
Out[179]:
Quantity
Date Buyer
2013-01-31 Carl 1
Mark 3
2013-10-31 Carl 6
Joe 9
2013-12-31 Carl 3
Joe 9
You have an ambiguous specification in that you have a named index and a column that could be potential groupers.
In [180]: df = df.set_index('Date')
In [181]: df['Date'] = df.index + pd.offsets.MonthEnd(2)
In [182]: df.groupby([pd.Grouper(freq='6M', key='Date'), 'Buyer']).sum()
Out[182]:
Quantity
Date Buyer
2013-02-28 Carl 1
Mark 3
2014-02-28 Carl 9
Joe 18
In [183]: df.groupby([pd.Grouper(freq='6M', level='Date'), 'Buyer']).sum()
Out[183]:
Quantity
Date Buyer
2013-01-31 Carl 1
Mark 3
2014-01-31 Carl 9
Joe 18
Taking the first rows of each group
Just like for a DataFrame or Series you can call head and tail on a groupby:
In [184]: df = pd.DataFrame([[1, 2], [1, 4], [5, 6]], columns=['A', 'B'])
In [185]: df
Out[185]:
A B
0 1 2
1 1 4
2 5 6
In [186]: g = df.groupby('A')
In [187]: g.head(1)
Out[187]:
A B
0 1 2
2 5 6
In [188]: g.tail(1)
Out[188]:
A B
1 1 4
2 5 6
This shows the first or last n rows from each group.
Taking the nth row of each group
To select from a DataFrame or Series the nth item, use nth(). This is a reduction method, and will return a single row (or no row) per group if you pass an int for n:
In [189]: df = pd.DataFrame([[1, np.nan], [1, 4], [5, 6]], columns=['A', 'B'])
In [190]: g = df.groupby('A')
In [191]: g.nth(0)
Out[191]:
B
A
1 NaN
5 6.0
In [192]: g.nth(-1)
Out[192]:
B
A
1 4.0
5 6.0
In [193]: g.nth(1)
Out[193]:
B
A
1 4.0
If you want to select the nth not-null item, use the dropna kwarg. For a DataFrame this should be either 'any' or 'all' just like you would pass to dropna:
# nth(0) is the same as g.first()
In [194]: g.nth(0, dropna='any')
Out[194]:
B
A
1 4.0
5 6.0
In [195]: g.first()
Out[195]:
B
A
1 4.0
5 6.0
# nth(-1) is the same as g.last()
In [196]: g.nth(-1, dropna='any') # NaNs denote group exhausted when using dropna
Out[196]:
B
A
1 4.0
5 6.0
In [197]: g.last()
Out[197]:
B
A
1 4.0
5 6.0
In [198]: g.B.nth(0, dropna='all')
Out[198]:
A
1 4.0
5 6.0
Name: B, dtype: float64
As with other methods, passing as_index=False, will achieve a filtration, which returns the grouped row.
In [199]: df = pd.DataFrame([[1, np.nan], [1, 4], [5, 6]], columns=['A', 'B'])
In [200]: g = df.groupby('A', as_index=False)
In [201]: g.nth(0)
Out[201]:
A B
0 1 NaN
2 5 6.0
In [202]: g.nth(-1)
Out[202]:
A B
1 1 4.0
2 5 6.0
You can also select multiple rows from each group by specifying multiple nth values as a list of ints.
In [203]: business_dates = pd.date_range(start='4/1/2014', end='6/30/2014', freq='B')
In [204]: df = pd.DataFrame(1, index=business_dates, columns=['a', 'b'])
# get the first, 4th, and last date index for each month
In [205]: df.groupby([df.index.year, df.index.month]).nth([0, 3, -1])
Out[205]:
a b
2014 4 1 1
4 1 1
4 1 1
5 1 1
5 1 1
5 1 1
6 1 1
6 1 1
6 1 1
Enumerate group items
To see the order in which each row appears within its group, use the cumcount method:
In [206]: dfg = pd.DataFrame(list('aaabba'), columns=['A'])
In [207]: dfg
Out[207]:
A
0 a
1 a
2 a
3 b
4 b
5 a
In [208]: dfg.groupby('A').cumcount()
Out[208]:
0 0
1 1
2 2
3 0
4 1
5 3
dtype: int64
In [209]: dfg.groupby('A').cumcount(ascending=False)
Out[209]:
0 3
1 2
2 1
3 1
4 0
5 0
dtype: int64
Enumerate groups
To see the ordering of the groups (as opposed to the order of rows within a group given by cumcount) you can use ngroup().
Note that the numbers given to the groups match the order in which the groups would be seen when iterating over the groupby object, not the order they are first observed.
In [210]: dfg = pd.DataFrame(list('aaabba'), columns=['A'])
In [211]: dfg
Out[211]:
A
0 a
1 a
2 a
3 b
4 b
5 a
In [212]: dfg.groupby('A').ngroup()
Out[212]:
0 0
1 0
2 0
3 1
4 1
5 0
dtype: int64
In [213]: dfg.groupby('A').ngroup(ascending=False)
Out[213]:
0 1
1 1
2 1
3 0
4 0
5 1
dtype: int64
Plotting
Groupby also works with some plotting methods. For example, suppose we suspect that some features in a DataFrame may differ by group, in this case, the values in column 1 where the group is “B” are 3 higher on average.
In [214]: np.random.seed(1234)
In [215]: df = pd.DataFrame(np.random.randn(50, 2))
In [216]: df['g'] = np.random.choice(['A', 'B'], size=50)
In [217]: df.loc[df['g'] == 'B', 1] += 3
We can easily visualize this with a boxplot:
In [218]: df.groupby('g').boxplot()
Out[218]:
A AxesSubplot(0.1,0.15;0.363636x0.75)
B AxesSubplot(0.536364,0.15;0.363636x0.75)
dtype: object
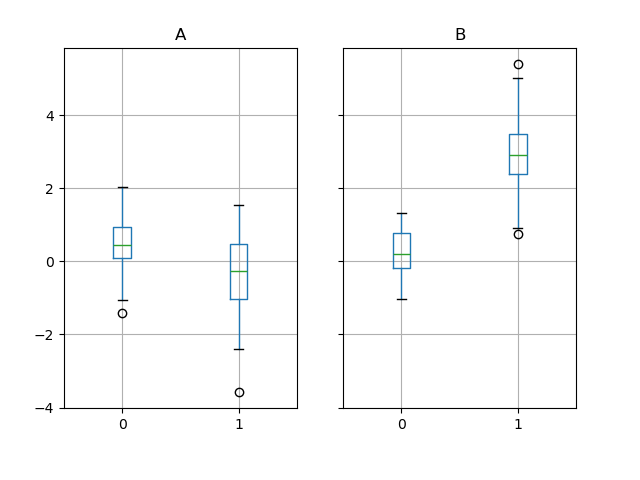
The result of calling boxplot is a dictionary whose keys are the values of our grouping column g (“A” and “B”). The values of the resulting dictionary can be controlled by the return_type keyword of boxplot. See the visualization documentation for more.
Warning
For historical reasons, df.groupby("g").boxplot() is not equivalent to df.boxplot(by="g"). See here for an explanation.
Piping function calls
Similar to the functionality provided by DataFrame and Series, functions that take GroupBy objects can be chained together using a pipe method to allow for a cleaner, more readable syntax. To read about .pipe in general terms, see here.
Combining .groupby and .pipe is often useful when you need to reuse GroupBy objects.
As an example, imagine having a DataFrame with columns for stores, products, revenue and quantity sold. We’d like to do a groupwise calculation of prices (i.e. revenue/quantity) per store and per product. We could do this in a multi-step operation, but expressing it in terms of piping can make the code more readable. First we set the data:
In [219]: n = 1000
In [220]: df = pd.DataFrame({'Store': np.random.choice(['Store_1', 'Store_2'], n),
.....: 'Product': np.random.choice(['Product_1',
.....: 'Product_2'], n),
.....: 'Revenue': (np.random.random(n) * 50 + 10).round(2),
.....: 'Quantity': np.random.randint(1, 10, size=n)})
.....:
In [221]: df.head(2)
Out[221]:
Store Product Revenue Quantity
0 Store_2 Product_1 26.12 1
1 Store_2 Product_1 28.86 1
Now, to find prices per store/product, we can simply do:
In [222]: (df.groupby(['Store', 'Product'])
.....: .pipe(lambda grp: grp.Revenue.sum() / grp.Quantity.sum())
.....: .unstack().round(2))
.....:
Out[222]:
Product Product_1 Product_2
Store
Store_1 6.82 7.05
Store_2 6.30 6.64
Piping can also be expressive when you want to deliver a grouped object to some arbitrary function, for example:
In [223]: def mean(groupby):
.....: return groupby.mean()
.....:
In [224]: df.groupby(['Store', 'Product']).pipe(mean)
Out[224]:
Revenue Quantity
Store Product
Store_1 Product_1 34.622727 5.075758
Product_2 35.482815 5.029630
Store_2 Product_1 32.972837 5.237589
Product_2 34.684360 5.224000
where mean takes a GroupBy object and finds the mean of the Revenue and Quantity columns respectively for each Store-Product combination. The mean function can be any function that takes in a GroupBy object; the .pipe will pass the GroupBy object as a parameter into the function you specify.
Examples
Regrouping by factor
Regroup columns of a DataFrame according to their sum, and sum the aggregated ones.
In [225]: df = pd.DataFrame({'a': [1, 0, 0], 'b': [0, 1, 0],
.....: 'c': [1, 0, 0], 'd': [2, 3, 4]})
.....:
In [226]: df
Out[226]:
a b c d
0 1 0 1 2
1 0 1 0 3
2 0 0 0 4
In [227]: df.groupby(df.sum(), axis=1).sum()
Out[227]:
1 9
0 2 2
1 1 3
2 0 4
Multi-column factorization
By using ngroup(), we can extract information about the groups in a way similar to factorize() (as described further in the reshaping API) but which applies naturally to multiple columns of mixed type and different sources. This can be useful as an intermediate categorical-like step in processing, when the relationships between the group rows are more important than their content, or as input to an algorithm which only accepts the integer encoding. (For more information about support in pandas for full categorical data, see the Categorical introduction and the API documentation.)
In [228]: dfg = pd.DataFrame({"A": [1, 1, 2, 3, 2], "B": list("aaaba")})
In [229]: dfg
Out[229]:
A B
0 1 a
1 1 a
2 2 a
3 3 b
4 2 a
In [230]: dfg.groupby(["A", "B"]).ngroup()
Out[230]:
0 0
1 0
2 1
3 2
4 1
dtype: int64
In [231]: dfg.groupby(["A", [0, 0, 0, 1, 1]]).ngroup()
Out[231]:
0 0
1 0
2 1
3 3
4 2
dtype: int64
Groupby by indexer to ‘resample’ data
Resampling produces new hypothetical samples (resamples) from already existing observed data or from a model that generates data. These new samples are similar to the pre-existing samples.
In order to resample to work on indices that are non-datetimelike, the following procedure can be utilized.
In the following examples, df.index // 5 returns a binary array which is used to determine what gets selected for the groupby operation.
Note
The below example shows how we can downsample by consolidation of samples into fewer samples. Here by using df.index // 5, we are aggregating the samples in bins. By applying std() function, we aggregate the information contained in many samples into a small subset of values which is their standard deviation thereby reducing the number of samples.
In [232]: df = pd.DataFrame(np.random.randn(10, 2))
In [233]: df
Out[233]:
0 1
0 -0.793893 0.321153
1 0.342250 1.618906
2 -0.975807 1.918201
3 -0.810847 -1.405919
4 -1.977759 0.461659
5 0.730057 -1.316938
6 -0.751328 0.528290
7 -0.257759 -1.081009
8 0.505895 -1.701948
9 -1.006349 0.020208
In [234]: df.index // 5
Out[234]: Int64Index([0, 0, 0, 0, 0, 1, 1, 1, 1, 1], dtype='int64')
In [235]: df.groupby(df.index // 5).std()
Out[235]:
0 1
0 0.823647 1.312912
1 0.760109 0.942941
Returning a Series to propagate names
Group DataFrame columns, compute a set of metrics and return a named Series. The Series name is used as the name for the column index. This is especially useful in conjunction with reshaping operations such as stacking in which the column index name will be used as the name of the inserted column:
In [236]: df = pd.DataFrame({'a': [0, 0, 0, 0, 1, 1, 1, 1, 2, 2, 2, 2],
.....: 'b': [0, 0, 1, 1, 0, 0, 1, 1, 0, 0, 1, 1],
.....: 'c': [1, 0, 1, 0, 1, 0, 1, 0, 1, 0, 1, 0],
.....: 'd': [0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1]})
.....:
In [237]: def compute_metrics(x):
.....: result = {'b_sum': x['b'].sum(), 'c_mean': x['c'].mean()}
.....: return pd.Series(result, name='metrics')
.....:
In [238]: result = df.groupby('a').apply(compute_metrics)
In [239]: result
Out[239]:
metrics b_sum c_mean
a
0 2.0 0.5
1 2.0 0.5
2 2.0 0.5
In [240]: result.stack()
Out[240]:
a metrics
0 b_sum 2.0
c_mean 0.5
1 b_sum 2.0
c_mean 0.5
2 b_sum 2.0
c_mean 0.5
dtype: float64

No comments:
Post a Comment